Bacterial Exchange
Categories: Journal no. 38, Diseases, Uganda, Bwindi, Gorilla Journal
The nature and frequency of human contact with wild primates is changing as a result of hunting, human encroachment on wildlife habitats, research, ecotourism, and other activities that bring people and primates into direct contact or close proximity (Adams et al. 2001). Such interactions may increase the risks of anthroponotic and zoonotic pathogen transmission, which can reduce human health as well as the health and viability of wild primate populations (Wallis & Lee 1999). Apes may be particularly susceptible to exchanging pathogens with people because they range widely into human habitats, are hunted and typically surrounded by high human-population densities. Additionally, many groups of free-ranging mountain gorillas (Gorilla beringei beringei) and chimpanzees have been habituated to humans for purposes of research and ecotourism, which brings them into close proximity to people on a regular basis.
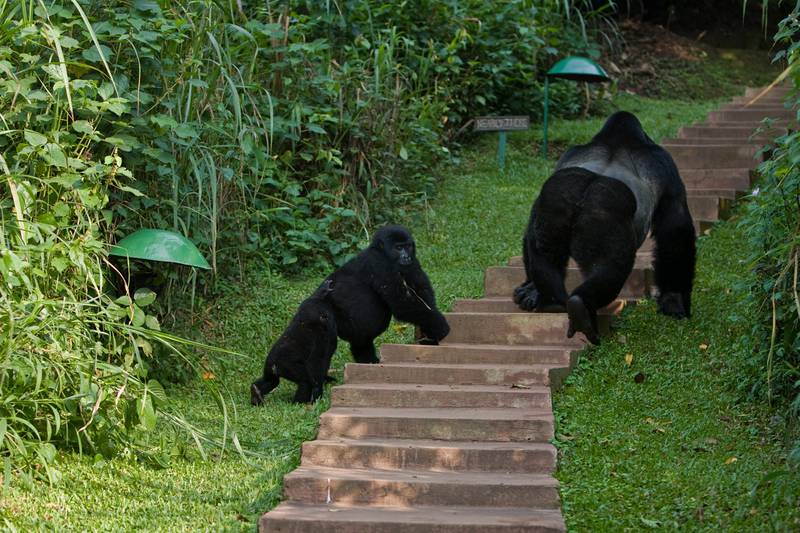
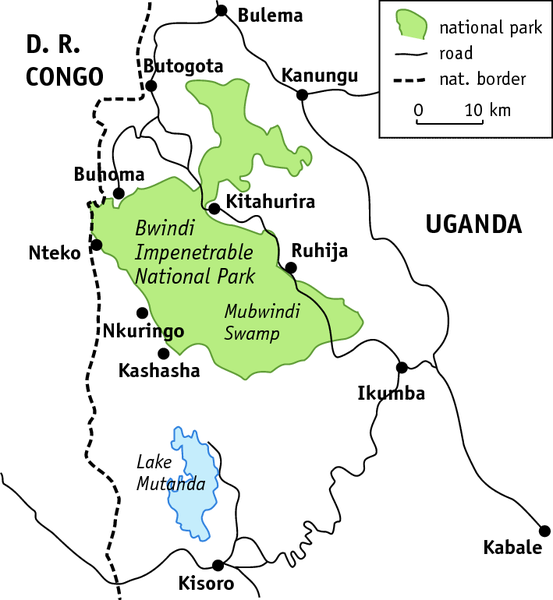
The study was carried out in Bwindi Impenetrable National Park to investigate whether habitat overlap influences rates and patterns of transmission of environmentally persistent and indirectly transmitted microbes between humans and wild apes. Mountain gorillas, an endangered taxon experiencing frequent contact with people and their livestock (goats, sheep, and cattle), were the main focus of the study. Three groups of mountain gorillas were targeted: Nkuringo, a group of 19 individuals that has been the focus of a tourism venture since 2004 and spends more than 67% of its time outside the park boundary; Kyagurilo, a group of 16 individuals that has been studied continuously for approximately 15 years by researchers but that is not visited by tourists; and a wild, unhabituated gorilla group that has no regular contact with humans and is not the subject of research. The population size of the wild gorilla group is unknown, but it is estimated at approximately 6 individuals on the basis of nest counts that were made at the time of sampling. The study also focused on people who interact with the mountain gorillas at high frequency as research workers or tour guides or because gorillas raid crops on their land.
Using a common gastrointestinal bacterium (Escherichia coli) as a model system, the nature of bacterial transmission across ape populations as a function of habitat overlap with people and livestock was investigated. Fecal samples from human volunteers, their livestock, and mountain gorillas were collected from May to August 2005 and bacteria were isolated and confirmed using standard microbiological methods. Genetic work was further done using previously described protocols. The susceptibility of the isolated bacteria to 11 antibiotics readily available to people in and around Bwindi Impenetrable National Park was measured.
Humans and livestock harboured bacteria that were very closely related to each other. Bacteria from all the three groups of gorillas were more closely related to bacteria from people employed in gorilla research and tourism than to bacteria from people in local villages. Across gorilla groups, genetic similarity between bacteria isolated from gorillas and those isolated from human populations was highest for the tourism group (group with highest human contact), lower for the research group (intermediate human contact), and lowest for the wild group (lowest or no human contact). Gorillas from the same group tended to share genetically similar bacteria. However, people working with the same gorilla group did not necessarily share genetically similar bacteria more than would be expected by chance alone.
Thirty-five percent of bacterial isolates from humans, 27% of isolates from livestock, and 17% of isolates from gorillas were clinically resistant to at least one of the antibiotics tested. Multiple resistances to Chloramphenicol, Streptomycin, Trimethoprim-sulfaxazole, and Tetracycline was observed in 4.2% of genetically distinct isolates, and multiple resistance to Ampicillin, Trimethoprim-sulfathaxazole, and Tetracycline was also observed in 7.2% of all genetically distinct isolates. This same pattern was observed in 20.3% of isolates from humans involved in gorilla work and 11.2% of genetically distinct isolates from humans from the village.
This means that habitat overlap among humans, livestock, and mountain gorillas can influence patterns of gastrointestinal bacterial exchange among species. Overall, gorilla populations that overlap in their use of habitat with people and livestock tend to harbor E. coli bacteria that are genetically similar to E. coli from those people or livestock. E. coli from the Nkuringo (tourism) gorilla group in particular were consistently most genetically similar to E. coli from local people and livestock. Mountain gorillas in the Nkuringo group spend a large percentage of their time outside the park boundary venturing into areas used by humans (Rwego 2004) and thus come into direct or indirect contact with villagers and their livestock. Conversely, gorillas in Kyagurilo group interact with the field assistants working with the group but not with local villagers, and gorillas from the wild group would rarely contact people or their habitats. These significant effects underscore that frequent contact and shared habitats, even on very fine scales, can influence bacterial transmission rates within and among populations of humans, apes, and livestock.
Antibiotic resistance was high in humans in this study. In rural Uganda, antibiotics are easily obtained over the counter and may be used indiscriminately. Antibiotics are rarely used for livestock in the Bwindi area, and administration of antibiotics to gorillas has been exceptionally rare. The presence of clinically resistant bacteria in gorillas (especially isolates resistant to multiple antibiotics) highly suggests that antibiotic-resistant bacteria are spreading from humans into the gorilla population. Such transmission appears to occur even between humans and gorilla groups that do not overlap with humans, although at a low rate, as evidenced by the presence of an isolate resistant to multiple antibiotics in the wild gorilla group. Local antibiotic use by humans seems to be responsible for the trends observed. Nearly the same patterns of antibiotic resistance were found in E. coli from humans and chimpanzees in a study carried out by Goldberg et al. (2007) in Kibale National Park, Uganda (approximately 200 km north of Bwindi and separated by a densely populated agricultural landscape).
These results should however be interpreted cautiously with respect to transmission. Genetic similarity between bacterial populations does not necessarily imply transmission in the conventional sense (i.e., direct exchange of microbes through direct or immediate contact). Transmission in the Bwindi system may occur indirectly and over extended time periods, perhaps through contaminated environmental sources such as soil and water. Goldberg et al. (2007) showed that bacterial gene flow was higher between chimpanzees and humans employed in chimpanzee research and tourism than between chimpanzees and people from local villages who rarely, if ever, share habitats with chimpanzees. This previous study also documented surprisingly high levels of antibiotic resistance in local people and the diffusion of antibiotic resistance to apes. Like chimpanzees, gorillas that are the subjects of research and tourism appear to be at increased risk of exchanging gastrointestinal microbes with people.
Overall, the patterns of genetic similarity and antibiotic resistance seen in the current study reflect the degrees to which apes, humans, and livestock interact. Habituation of mountain gorillas to humans for the purposes of research and tourism also appears to be associated with increased risks of gastrointestinal bacterial transmission between the species. Concerns about pathogen transmission already underlie many of the regulations in place governing interactions between people and apes (e.g. minimum observational distances, maximum observation times). These results suggest, however, that apes even in well-managed situations may be at increased risk of pathogen exchange with humans and livestock. If common sources of environmental contamination underlie the trends that have been documented, then preventing direct or even close contact between people and mountain gorillas may not be sufficient for preventing microbial exchange. This conclusion may apply to gastrointestinal pathogens and to pathogens transmitted by other modes, such as through the respiratory system, that represent serious and potentially epidemic disease threats to wild apes. Strategies such as discouraging people from defecating in the forest, encouraging hand washing before and after entering the forest, mandating the wearing of aerosol-limiting face masks for people entering ape habitats, and encouraging employee health programs would be reasonable strategies to limit bacterial exchange between people and apes, which would safeguard ape health and aid conservation efforts.
Innocent B. Rwego, Thomas R. Gillespie, Gilbert Isabirye-Basuta and Tony L. Goldberg
We thank Uganda Wildlife Authority and Uganda National Council for Science and Technology for granting us permission to conduct this study. Wildlife Conservation Society Research Fellowship and Morris Animal Foundation supported this study.
The full scientific report (from which this article is taken) is available from Conservation Biology 22 (6), 1600-1607.
References
Adams, H. R. et al. (2001) Self-reported medical history survey of humans as a measure of health risks to chimpanzees (Pan troglodytes schweinfurthii) of Kibale National Park. Oryx 35, 308-312
Goldberg, T. L. et al. (2007) Patterns of gastrointestinal bacterial exchange between chimpanzees and humans involved in research and tourism in western Uganda. Biological conservation 135, 511-517
Rwego, I. B. (2004) Prevalence of Clinical Signs in Mountain gorillas, Bwindi Impenetrable National Park. M.Sc dissertation, Department of Wildlife and Animal Resources Management, Makerere University, Kampala
Wallis, J. & Lee, D. R. (1999) Primate Conservation: the prevention of disease transmission. International Journal of Primatology 20, 803-826